The Hidden Problem in Modern Biology
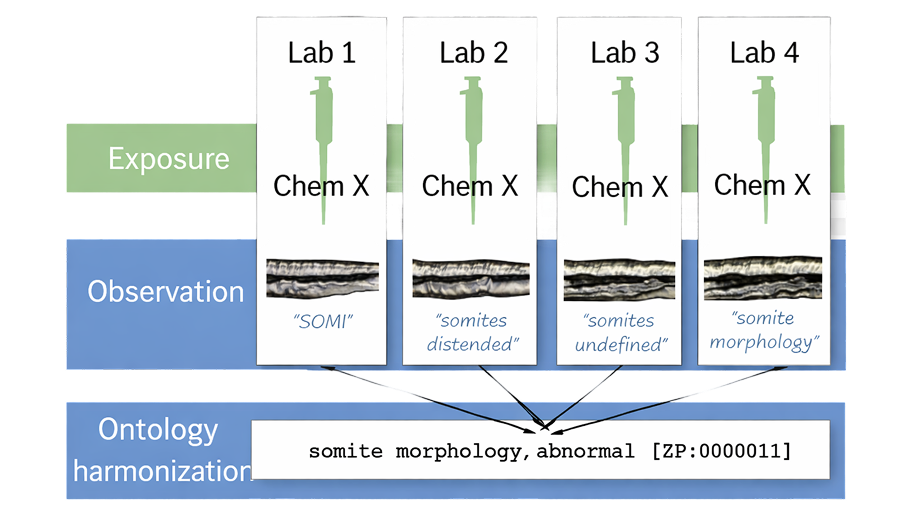
If you have ever tried to compare results across zebrafish studies, you have likely encountered a frustrating issue. One paper reports “reduced swimming,” another describes “hypoactivity,” and a third refers to “decreased locomotor response.” It is not always clear whether these observations are identical, similar, or meaningfully different.
When this inconsistency is multiplied across hundreds of studies, laboratories, and regulatory submissions, it becomes a major barrier. Comparing findings, integrating datasets, and reusing data all become unnecessarily difficult. This is one of the central bottlenecks in modern biology and exactly the kind of problem ontologies are designed to solve.
What is an Ontology?
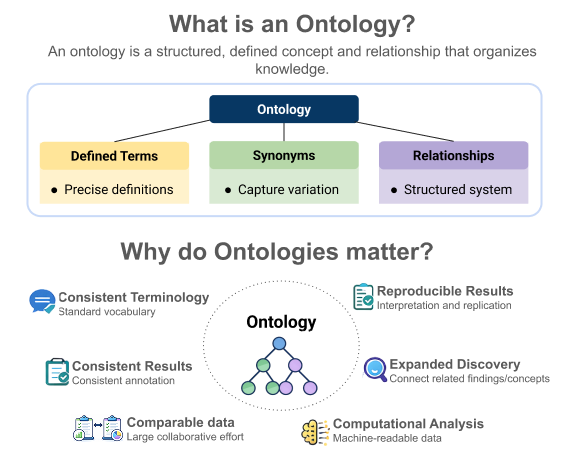
At its core, an ontology is a standardized way of naming concepts and describing how those concepts are related to one another. It goes beyond a simple glossary by not only defining terms but also organizing them in a structured, connected framework.
An ontology typically includes:
- Clearly defined terms (e.g., “heart,” “liver,” or “brain”) along with precise definitions
- Synonyms to account for variation in terminology
- Explicit relationships between terms (e.g., “is a” or “part of”) which help organize knowledge into a connected system
This structure allows both researchers and software to understand how concepts fit together.
Why Ontologies Matter
As biology becomes increasingly data-driven, ontologies are becoming essential infrastructure. They directly impact how research is conducted, shared, and reused.
Ontologies standardize language. Different labs often use different terms for the same concept. Ontologies provide a shared vocabulary with precise definitions, reducing ambiguity.
Ontologies make data comparable across studies. With an ontology, data can be aligned more easily, enabling cross-study analysis, meta-analysis, and large collaborative efforts.
Ontologies improve search and discovery. Instead of relying on exact keyword matches, ontology-based systems can retrieve related concepts automatically.
Ontologies support reproducibility by reducing ambiguity in how results are described, making experiments easier to interpret and replicate through consistent, well-defined terminology.
Ontologies enable modern computational approaches, including machine learning and artificial intelligence. These methods depend on structured, machine-readable data, and ontologies provide exactly that foundation.
Why Ontologies Matter for the Zebrafish Community
In zebrafish research, ontologies might sound like a technical concept, but they solve a very practical problem: everyone describes things a little differently. One lab might record a behavior as “reduced swimming,” while another calls it “hypoactivity.” Ontologies provide a shared, standardized vocabulary with clear definitions, so researchers are essentially speaking the same language. This makes a big difference when trying to compare results across studies, especially in a field like zebrafish research where experiments often generate large, complex datasets. With consistent terminology, data from different labs can be combined more easily, enabling broader analyses and collaborations.
They also make it much easier to find relevant information. Instead of relying on exact keyword matches, ontology-based systems can connect related ideas. So, for example, if you search for “heart defects”, an ontology-based system can also surface studies describing related findings such as “cardiac malformations” or “cardiac abnormality”, even if those exact words were not used in your original search. Another benefit is reproducibility: when terms are clearly defined and used consistently, it becomes easier for others to understand exactly what was observed and compared to and to replicate the experiment.
Ontologies also open the door to modern computational approaches, such as machine learning and artificial intelligence. These methods rely on data that is not only digital, but structured and consistently defined so that computers can interpret it correctly. Ontologies provide this foundation by organizing information into clearly defined concepts and relationships, making datasets machine-readable and interoperable. As a result, researchers can apply advanced analytical tools to automatically detect patterns, compare results across large datasets, and even generate new hypotheses. In zebrafish research, this means being able to extract deeper insights from complex phenotypic and behavioral data, ultimately accelerating discovery and improving the predictive power of scientific studies.
The benefits extend beyond research into regulatory decision-making because zebrafish are widely used in chemical safety and toxicity testing by organizations such as the U.S. Environmental Protection Agency and the Organisation for Economic Co-operation and Development. Regulators rely on clear, comparable data to assess risk and make informed judgments. Ontologies support this process by standardizing how study results are described across different submissions. This leads to greater transparency, more efficient review processes, and better integration of evidence from both academic and industry sources. Ultimately, using ontologies helps ensure that risk assessments are more consistent, reliable, and defensible.
Examples of Ontologies
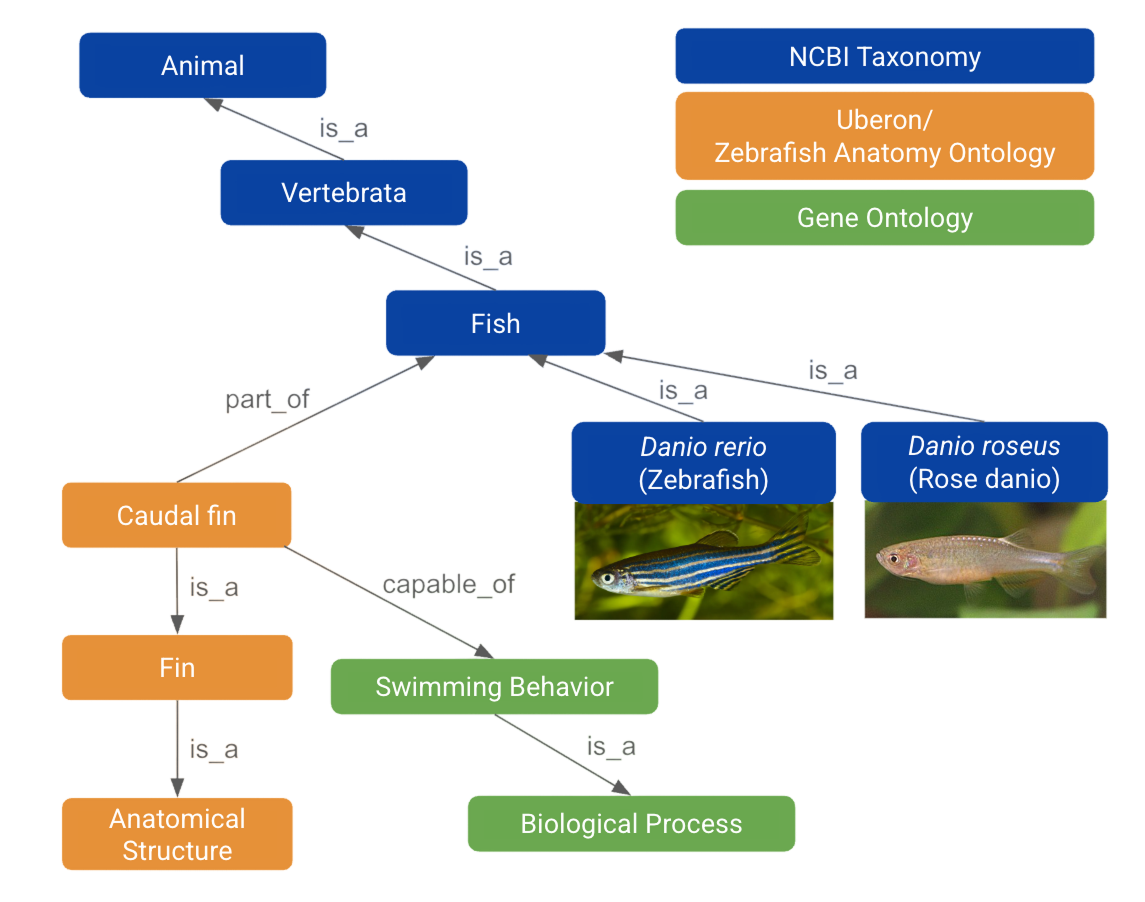
Anatomical ontologies
Phenotype ontologies
- Zebrafish Phenotype Ontology (ZP)
- Unified Phenotype Ontology (UPheno)
- Phenotype and Trait Ontology (PATO)
Exposure and toxicology ontologies
- Exposure Ontology (ExO)
- Environment Exposure Ontology (ECTO)
- Chemical Entities of Biological Interest (ChEBI)
Key Takeaway
Without ontologies, valuable data often remains siloed, inconsistently described, and difficult to reuse and/or compare. This limits the impact of research and slows scientific progress.
Ontologies are not just a technical detail or a niche informatics tool. They are essential infrastructure for modern science. By enabling a shared language, supporting data integration, and powering computational analysis, ontologies help unlock the full value of biological data.
